Labeled extract items should be used to describe the event that transformed an extract material in a labeled extract material. Labeled extracts can be created from extract items or from one or more labeled extract items. When a labeled extract is created from several other labeled extracts, a pooling event is performed.
During the transformation from extracts to labeled extracts, it is possible to provide information about the protocol used to perform this task. It is not enforced in BASE but it should serve as guidance when devising the granularity of the extract processing task. Also, it is good practice to provide protocol information.
Beside the regular way of using the button in → , a labeled extract can be created in one of following ways.
- pooling selected labeled extracts
-
The toolbar at the list page of labeled extracts contains an addition button. This button allows users to create pooled labeled extracts by selecting the list of labeled extracts used to derived a new labeled extract and then click on the button.
This provides an easy and simple way to create pooled labeled extracts. The result of such process is the creation of a new labeled extract, in which, when navigating to the parent tab, shows that all the labeled extracts involved are already set and listed in the Labeled Extract box of the tab.
- from the extract pages.
Following the laboratory workflow, a natural way to create a labeled extract from an extract is to click on the
 from the labeled extract column of the extract list view. Corresponding
button,
is located on the single item view page for each extract. By creating a
labeled extract from an extract page will automatically set the extract
as a parent.
from the labeled extract column of the extract list view. Corresponding
button,
is located on the single item view page for each extract. By creating a
labeled extract from an extract page will automatically set the extract
as a parent.
- from single item view of a label
Click on the to use the current label and create a new labeled extract with it pre-selected as Label .
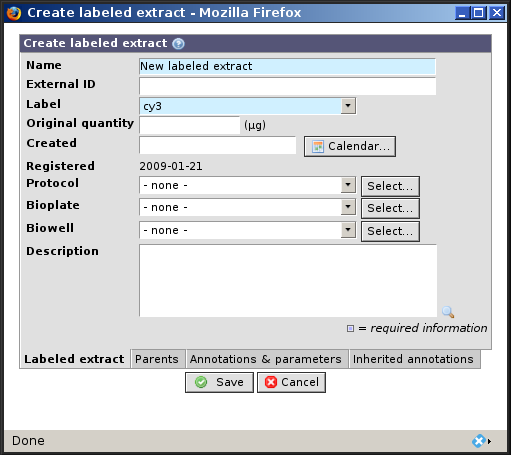
- Name
The name of the labeled extract (required). BASE by default assigns names to labeled extract(by suffixing
lbe#when creating a labeled extract from an existing extract orNew labeled extractotherwise) but it is possible to edit it at will- Label
Used to specify which dye or marker was used in the labeling reaction (required).
- External ID
An id used to recognize the labeled extract outside BASE.
- Original quantity
The mass of labeled extract that was created.
- Created
A date should be provided. The information can be important when running quality controls on data and account for potential confounding factor (e.g. day effect).
- Registered
This is automatically populated with a date at which the labeled extract was actually entered in BASE system.
- Protocol
The labeling protocol that was used to produce the labeled extract.
- Bioplate
The bioplate where this labeled extract is located.
- Biowell
Biowell that holds this labeled extract. Bioplate has to be defined before biowell can be selected.
- Description
A free text field to report any information that can not be captured otherwise.
This important tab allows users to select the labeled extract origin. BASE distinguished between two cases which are controlled by the Pooled radio-button.
-
The parent is an extract
The radio-button is set to No . The Extract select button is active and allows users to point to one and only one extract from which the labeled extract originates from.
-
The parent is another labeled extract
The radio-button has to be set to Yes . Upon selection, the extract select button is deactivated and the labeled extracts box and button are activated. This allows users to specify one or more extracts to be selected from the labeled extract list view page.
As seen in the biosource and sample sections, this tab allows users to further supply information about the labeled extract provided they have defined annotation types to annotate labeled extract items or have such elements shared to them.
![[Important]](../../gfx/admonitions/important.gif) |
Important |
|---|---|
In order to use this feature, annotation type must be declared and made available. To learn more about annotation types and how these are set, please refer to Chapter 11, Annotations . |
This tab contains a list of those annotations that are inherited from the parents of the labeled extract. Information about dealing with inherited annotations can be found in Section 11.4.1, “Inheriting annotations from other items” .