A hybridization event corresponds to the application of one or more labeled extracts materials to a microarray slide under conditions detailed in hybridization protocols. Use → to get to the hybridizations.
In BASE, there are two possible routes to create an hybridization object except the common way with the button at hybridization list page.
- from the labeled extract list view page
-
Select at least one labeled extract, to create a hybridization from, by ticking the selection boxes before the name field.
Click on the from the toolbar of labeled extract list view.
- from a labeled extract single item page
When viewing a label extract in single item view, click on the button from the toolbar of the labeled extract item view.
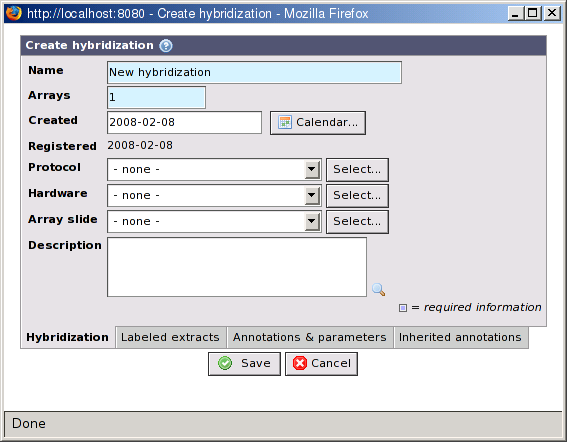
- Name
New hybridizationis the BASE default name but it is strongly advise to provide a meaningful and unique name (required).- Arrays
The number of sub-arrays on the slide that was used in this hybridization. The default value is 1, but some platforms, for example Illumina, has slides with 6 or 8 arrays. When the array-slide property below is changed this value will be updated automatically to be consistent with the number of sub-arrays on the used array-design.
- Created
A date should be provided. The information can be important when running quality controls on data and account for potential confounding factor (e.g. to account for a day effect)
- Registered
This field is automatically populated with a date at which the hybridization was entered in BASE system.
- Protocol
The hybridization protocol that was used to do the hybridization.
- Hardware
The hybridization-station that was used during the hybridization.
- Array slide
-
The array slide that was used in the hybridization.
![[Note]](../../gfx/admonitions/note.gif)
Note Ideally, The Array Slides should have been created but for those users with permission to do, Array Slides could be generated at that point.
- Description
A free text field to report any information that can not be captured otherwise
This important tab allows users to select the labeled extracts applied to an array slide, and specify the amount of material used, expressed in microgram.
Use the button to add items and the button to remove items. Select one or several labeled extracts in the list and write the used mass and sub-array index in the fields below.
As seen in the biosource and sample sections, this tab allows users to supply further information about the hybridization provided annotation types have been defined or shared to annotate hybridization items.
![[Important]](../../gfx/admonitions/important.gif) |
Important |
|---|---|
In order to use this feature, annotation type must be declared and made available. To learn more about annotation types and how these are set, please refer to Chapter 11, Annotations |
This tab contains a list of those annotations that are inherited from the labeled extracts. Information about dealing with inherited annotations can be found in Section 11.4.1, “Inheriting annotations from other items” .