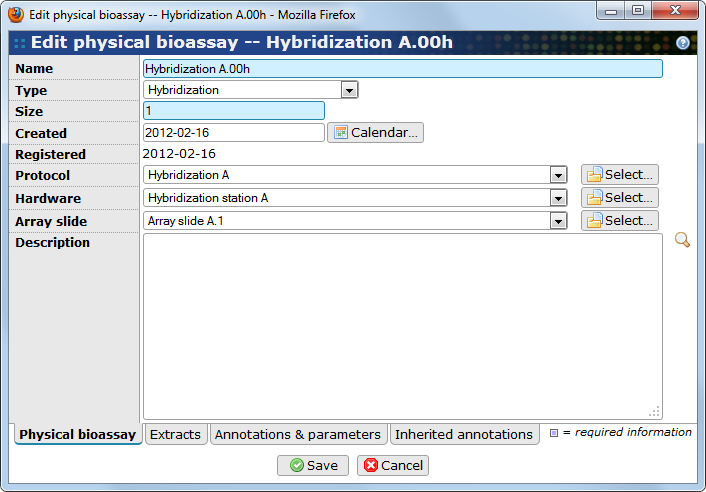
A physical bioassay represents the application of one or more extracts to an experimental setup designed to measure quantities that we are interested in. For example, a Hybridization event corresponds to the application of one or more Labeled extracts materials to a microarray slide under conditions detailed in hybridization protocols. Use to get to the bioassays.
In BASE, there are two possible routes to create a physical bioassay except the common way with the button at the list page.
- from the extract list view page
-
Select at least one extract, to create a bioassay from, by ticking the selection boxes before the name field. Click on the in the toolbar.
- from the extract single-item page
-
When viewing an extract in single-item view, click on the button in the toolbar.
- Name
-
The bioassay's name (required). It is recommended that the default name is replaced with something that is unique.
- Type
-
The subtype of the bioassay. The list may evolve depending on additions by the server administrator. Selecting the proper subtype is recommended and enables BASE to automatically guess the most likely subtype when assignint source biomaterials and when creating derived bioassays. See Chapter 12, Item subtypes for more information.
- Size
-
The size of the bioassay is the number of independent positions on the bioassay. Depending on the characteristics of the bioassay, multiple biomaterials may share the same position, but are then usually required to have different tags. Two biomaterials in different positions can use the same tag. The default value is 1, but some platforms, for example Illumina BeadArrays, has slides with 6 or 8 positions and sequencing flow cells have 8 lanes.
- Created
-
A date should be provided. The information can be important when running quality controls on data and account for potential confounding factor (e.g. to account for a day effect).
- Registered
-
This field is automatically populated with a date at which the hybridization was entered in BASE system.
- Protocol
-
The protocol that was used to create the bioassay.
- Hardware
-
Information about the machine (if any) that was used when creating the bioassay.
- Array slide
-
The array slide that was used for the bioassay.
- Description
-
A free text field to report any information that can not be captured otherwise.
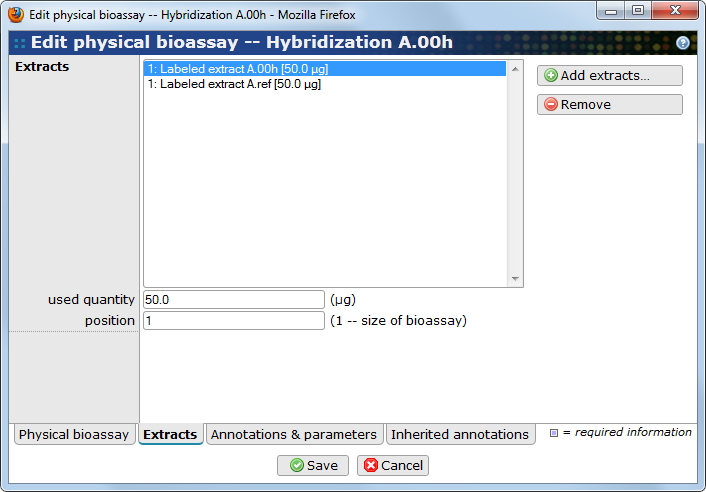
This important tab allows users to select the extracts used by the bioassay, and specify the amount of material used, expressed in microgram.
Use the button to add items and the button to remove items. Select one or several extracts in the list and write the used mass and position number in the fields below the list.
The Annotations tab allows BASE users to use annotation types to refine bioassay description. More about annotating items can be read in Section 10.2, “Annotating items”
This Inherited annotations tab contains a list of those annotations that are inherited from the bioassay's parents. Information about working with inherited annotations can be found in Section 10.2.2, “Inheriting annotations from other items”.