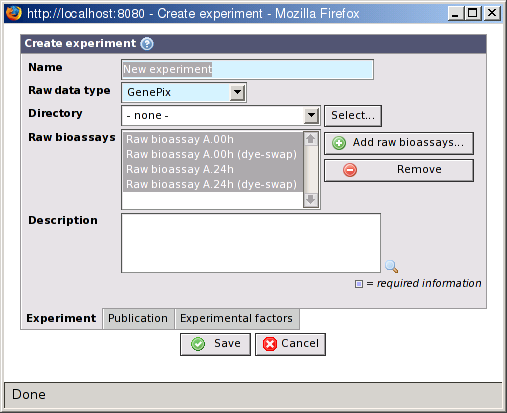
Experiments are the starting point for analysis. When you have uploaded and imported your raw data, collected and registered all information and annotations about samples, hybridizations, and other items, it is time to collect everything in an experiment.
To create a new experiment you can either mark one ore more raw biossays on the raw bioassays list view and use the button. You can also create a new experiment from the experiments list view.
- Name
The name of the experiment.
- Raw data type
The raw data type to use in the experiment. All raw bioassays must have raw data with this type.
- Directory
A directory in the BASE file system where plug-ins can save files that are generated during the analysis. This is optional and if not given the plug-ins must ask about a directory each time they need it. Use the button to browse the file system or create a new directory.
- Raw bioassays
The raw bioassays you want to analyze in this experiment. If you created the experiment from the raw bioassays list the selected raw bioassays are already filled in. Use the button to add more raw bioassays or the button to remove the selected raw bioassays from the list.
- Description
A description of the experiment.
Click on the button to save the changes or on to abort.
On this tab you can enter information about a publication that is the result of the experiment. All of this information is optional.
- PubMedId
The ID of the publication in the PubMed database.
- Title
The title of the publication.
- Publication date
The date the article was published. Use the button to select a date from a pop-up window.
- Abstract
The article abstract.
- Experiment design
An explanation of the experiment design.
- Experiment type
A description of the experiment type.
- Affiliations
Partners and other related organisations that have helped with the experiment.
- Authors
The list of authors of the publication.
- Publication
The body text of the publication.
Click on the button to save the changes or on to abort.
The experimental factors of an experiment are the variables you are studying in the experiment. Typically the value of an experimental factor is varied between samples or group of samples. Different treatment methods is an example of an experimental factor.
In the BASE world an experimental factor is the same as an annotation type. Since you probably have lots of annotations on your items that are not relevant for the experiment you must select the annotations types that should make up the experimental factors of the experiment.
Use the button to select the annotation types that should be used as experimental factors. The button removes the selected annotation types.
Click on the button to save the changes or on to abort.
To be able to use the values of the experimental factors in the analysis of your data the values must be accessible from the raw bioassays. Since most of your annotations are probably made at the sample or biosource level the raw bioassays must inherit those annotations. Read Section 11.4.1, “Inheriting annotations from other items” for more information about this.
![[Tip]](../../gfx/admonitions/tip.gif) |
Tip |
|---|---|
Use the Item overview function to verify that all your raw bioassays has been annotated or inherited values for all experimental factors. If not, you should do that before starting with the analysis. |
Tab2Mage format is a tab-delimited format veted by EBI's ArrayExpress repository for submission microarray data. Tab2Mage format has been chosen by BASE to provide an easy way for data deposition to public repository when submitting a manuscript and publishing experimental data.
BASE has been engineered to closely map the MIAME concepts and a number of BASE entities can be mapped directly to Tab2Mage elements. However, since MIAME is focused on microarray processing workflow, information about the biological sample is down to the user. To accommodate the annotation needs of users, BASE provides a mechanism that allows annotation customization to meet user specific requirements. BASE allows to create annotation type for quantitative annotation and qualitative annotation
BASE can export an experiment to Tab2Mage format thanks to a dedicated export plug-in. For the plug-in to work, it is important to understand that information recorded in BASE should be formatted following a small number of rules. Failing to do so may impair the possibility of exporting to ArrayExpress.
![[Note]](../../gfx/admonitions/note.gif) |
Note |
|---|---|
The Tab2Mage export plug-in has not yet been included in the main distribution. Hopefully, it will appear in the next (2.4) release. |
Tab2Mage specifications only allow BioSource items to be annotated with BioMaterialCharacteristics.
![[Warning]](../../gfx/admonitions/warning.gif) |
Warning |
|---|---|
All BASE Annotation Types used to annotate at the level of Sample and Extract items will be lost during the export in Tab2Mage format in order to comply with the ArrayExpress Tab2Mage parser. |
![[Note]](../../gfx/admonitions/note.gif) |
Note |
|---|---|
In the context of data exchange between BASE instances, the export function can be altered to allow attachment of annotations to items other than biosources, therefore avoiding loss of information. |
To associate units to BASE annotation types and remain compatible with Tab2Mage specifications, users need to adhere to the following convention:
annotation_type_name(unit_name) as in
body mass(kg) or concentration(mg/ml)
![[Warning]](../../gfx/admonitions/warning.gif) |
Warning |
|---|---|
Only one unit can be specified at any one time
for any given annotation type. In order to
enable Tab2Mage support, it might be necessary
to declare several related Annotation Type in
order to report similar kind of information but
expressed in a different unit. Specifying Age
for instance is a good example on how to deal
with such cases: One should create several
related annotation types e.g. |
In order to ensure MIAME compliance, Tab2Mage specifications cater for reporting parameters attached to protocols and all parameters attached to a protocol should be declared in the protocol section of a Tab2Mage file.
In BASE terms, Tab2Mage elements such as BioMaterialCharacteristics, Parameter or FactorValues are all annotation types. But, it is necessary to flag those annotations types meant to be used as protocol parameters as such so that they can identified by the Tab2Mage exporter and handled appropriately.
![[Warning]](../../gfx/admonitions/warning.gif) |
Warning |
|---|---|
It is not possible to use the same annotation type Temperature for reporting a patient body temperature (which is a Biomaterial Characteristic) and hybridization temperature (which is a protocol parameter). Hence it will be necessary to declare 2 distinct annotation types: |
Annotation type to be used as BioSource characteristics:
body temperature (degree_C)Annotation type to be used as protocol parameter:
hybridization temperature (degree_C)
It is a MIAME requirement to identify Experimental Variables when submitting data to ArrayExpress (provided the study is an intervention study). Therefore, BASE users willing to use the Tab2Mage export function will have to declare Experimental Factors using the the Experimental Factor tab available when editing experiments. See Section 18.3.2, “Experimental factors” for more information.
Values for the experimental factors are take from annotations. The annotation must exist at the raw bioassay level, which probably means that you have to inherit the annotation from some other item, for example, a biosource or a sample. It is also possible to use a protocol parameter as experimental factor. See Chapter 11, Annotations for more information about annotations.